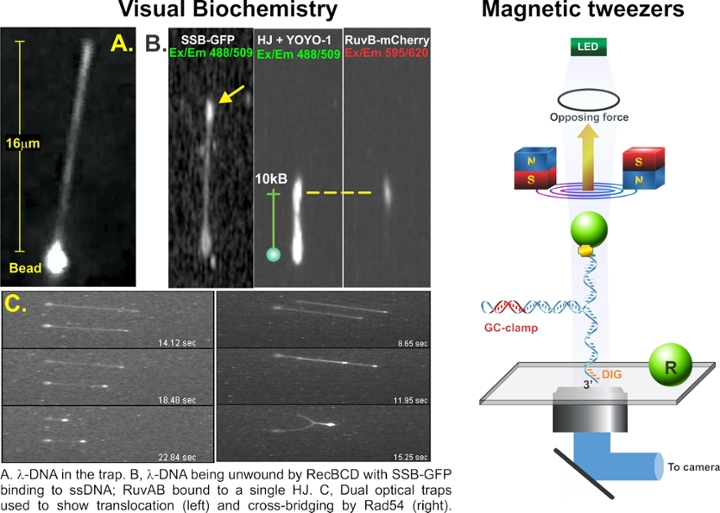
Two, complementary single-molecule approaches are used to provide insight into DNA transactions. The first is what we call “visual biochemistry”, a technique I pioneered that combines optical tweezers, microfluidics, and high-resolution fluorescence microscopy. Here, 1 micron polystyrene beads are bound to fluorescent-labeled DNA. When the beads are optically trapped, fluid flow past the bead stretches the DNA out like a rope behind a boat. When DNA binding proteins process the DNA, the molecule can shorten (DNA helicase unwinding) and fluorescent SSB protein can bind to the unwound stands of DNA. Single RuvAB complexes will bind to a Holliday Junction to facilitate branch migration or dsDNA translocases like Rad54 can either translocate or cross-bridge dsDNA to enhance homologous recombination. To visualize cross-bridging, dual optical traps were used to bring protein-coated DNA molecules together. The second technique known as magnetic tweezers is used to achieve single base-pair resolution. Here a DNA molecule is attached to a coverslip surface at one end and to a super-paramagnetic bead at the other. The bead is manipulated by the magnetic tweezers to other stretch or relax the DNA. We have manipulated single hairpin DNA molecules to understand fork regression by RecG or the mechanism of SSB displacement by the helicase.