
(From left: Dr. Reid, Jane Morwitzer, Enakshi Roy and Ope Oyewole.)
Alphavirus
Chikungunya virus (CHIKV) is an Old-World alphavirus that causes fever, rashes and arthralgia. It is an arboviral disease that is vectored by Aedes aegypti and Aedes albopictus and is endemic in several African and Asian countries. First isolated in Tanzania in 1953, the virus has since spread to Asia, Europe and the Americas, making it a re-emerging infectious disease. There is currently no vaccine or anti-viral therapy for CHIKV, thus a better understanding of mechanisms of infection as well as host involvement during infection is needed.
Our lab is interested in the role of host proteins during CHIKV infection. Previous work identified Sphingosine kinase 2 (SphK2) as an essential host protein during infection. Using mutational analysis, we have identified the viral interacting partner for SphK2 as nsP3. Current work is focused on identifying the specific viral and host motifs necessary for interaction.
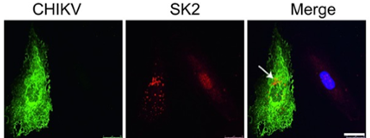
Figure 1. SK2 is re-localized during CHIKV infection. MCF10A cells were uninfected or infected with CHIKV at a MOI of 5 for 24 h. The viral E2 protein (green), SK2 (red), and Hoechst nuclear stain (blue) were visualized by confocal microscopy. Scale bar = 25 mm
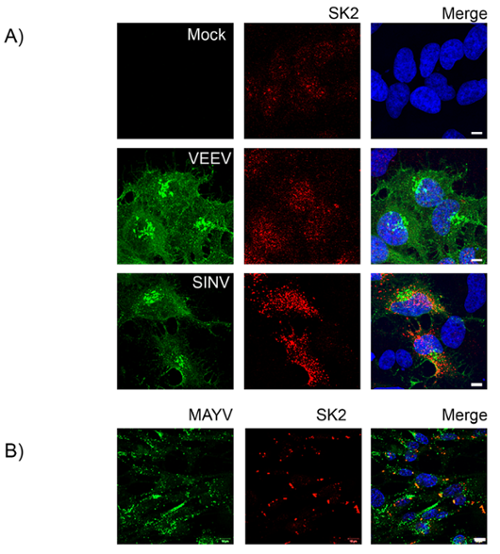
Figure 2. SK2 re-localizes during infection with arthritogenic, but not encephalitic alphaviruses. A) BHK21 cells were either mock infected or infected with SINV (MOI = 1) or VEEV (MOI = 0.01) (A) or MAYV (MOI = 5) (B) for 24 h. The cells were then fixed and stained for viral antigen (green), SK2 (red) and Hoechst nuclear stain (blue). Scale bars = 5 µm (A), 10 µm (B).
Joint/Cartilage Project
The chronic pathology of CHIKV infection has been likened to that of Rheumatoid Arthritis (RA) due to the evidence of bone pathology and inflammation. A better understanding of interactions in the bone microenvironment is needed to provide insight into how the virus is able to cause chronic sequelae in infected patients.
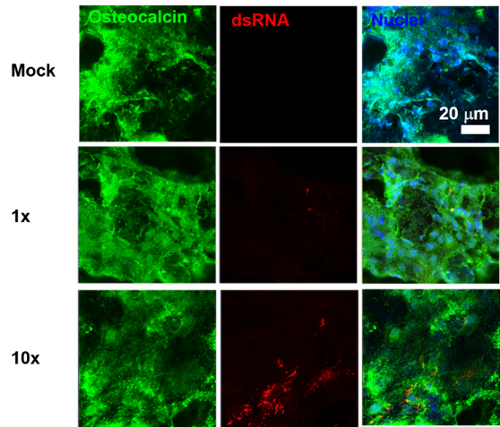
Figure 3. Infection of Osteoblasts in 3D culture. Osteoblasts were infected with CHIKV at a MOI 5 for 24 h. The cells were fixed and stained for viral antigen (red) osteocalcin (green) an osteoblast marker and Hoechst nuclear stain (blue)
 Filoviruses
Filoviruses
Filovirius are single-stranded negative-sense RNA viruses within the order Mononegavirales. These viruses include the genus ebola virus, marburgvirus and cuevavirus. Generally, these viruses are the causative agent of severe, often fatal, hemorrhagic fever in humans, with case fatality rates as high as 90%. The primary focus of our laboratory are ebolaviruses. Since its emergence in 1976, sporadic localized outbreaks have typically characterized these viruses, but the unprecedented spread of the 2013–2016 West African epidemic and its high death toll highlights the global public health threat that it poses. The recent 2017 and 2018 outbreaks in Democratic Republic of the Congo further emphasize the urgency for therapeutic interventions. Despite this, there remains no licensed therapeutics available, although a number of promising candidates are currently under investigation. Novel therapeutic targets will thus be useful to the continued development of effective countermeasures. (Above left image: Ebola Virus Particle courtesy of wikimedia.org)
Our laboratory primarily focuses on Ebola virus (EBOV formerly known as Zaire ebola virus). This species has been associated with the most devastating outbreaks and appears to be the most virulent. The primary focus of our work is to understand better the molecular mechanisms underlying EBOV pathogenesis. Towards this goal, we are interested in host-protein interactions during EBOV infection. Previous work identified interactions of EBOV nucleoprotein (NP) with host proteins RUVBL1 and RUVBL2. Future studies will involve determining the role of these proteins in EBOV biology. Additional studies are also underway to elucidate the role of additional candidate host interactors.
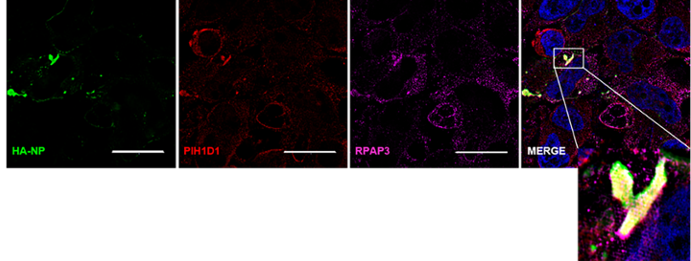
Figure 4. The R2TP complex components RPAP3 and PIH1D1 colocalize with HA-NP. HeLa cells were transfected with vector control or HA-NP. Twenty-four hours later, the cells were fixed and processed for the immunofluorescence detection of endogenous PIH1D1 or RPAP3 in the presence of vector control or HA-NP. Representative images of endogenous localization pattern of PIH1D1 and RPAP3 in the presence of HA-NP. HA-NP (green), RPAP3 (red), PIH1D1 (magenta), and Hoechst 33342 nuclear stain (blue) were visualized by SIM. Scale bars = 20 µM.

